Is Dna Read From 5 to 3
Arthur Kornberg compared Deoxyribonucleic acid to a tape recording of instructions that tin exist copied over and over. How do cells make these near-perfect copies, and does the process always vary?
Scientists take devoted decades of effort to agreement how deoxyribonucleic acrid (DNA) replicates itself. In unproblematic terms, replication involves apply of an existing strand of DNA as a template for the synthesis of a new, identical strand. American enzymologist and Nobel Prize winner Arthur Kornberg compared this procedure to a tape recording of instructions for performing a chore: "[E]xact copies can be made from it, every bit from a tape recording, so that this information tin be used once more and elsewhere in fourth dimension and space" (Kornberg, 1960).
In reality, the process of replication is far more circuitous than suggested past Kornberg's illustration. Researchers typically utilize elementary bacterial cells in their experiments, but they still do non have all the answers, particularly when information technology comes to eukaryotic replication. Nonetheless, scientists are familiar with the bones steps in the replication process, and they go on to rely on this data as the basis for continued research and experimentation.
The Molecular Machinery of Bacterial DNA Replication
A typical bacterial cell has anywhere from virtually 1 1000000 to 4 million base of operations pairs of DNA, compared to the 3 billion base pairs in the genome of the common firm mouse (Mus muscle). Nevertheless, fifty-fifty in bacteria, with their smaller genomes, DNA replication involves an incredibly sophisticated, highly coordinated series of molecular events. These events are divided into 4 major stages: initiation, unwinding, primer synthesis, and elongation.
Initiation and Unwinding
During initiation, so-called initiator proteins bind to the replication origin, a base-pair sequence of nucleotides known every bit oriC. This binding triggers events that unwind the Deoxyribonucleic acid double helix into ii single-stranded Deoxyribonucleic acid molecules. Several groups of proteins are involved in this unwinding (Figure one). For example, the DNA helicases are responsible for breaking the hydrogen bonds that join the complementary nucleotide bases to each other; these hydrogen bonds are an essential feature of James Watson and Francis Crick's three-dimensional DNA model. Because the newly unwound single strands take a tendency to rejoin, another group of proteins, the unmarried-strand-binding proteins, keep the single strands stable until elongation begins. A tertiary family of proteins, the topoisomerases, reduce some of the torsional strain acquired by the unwinding of the double helix.
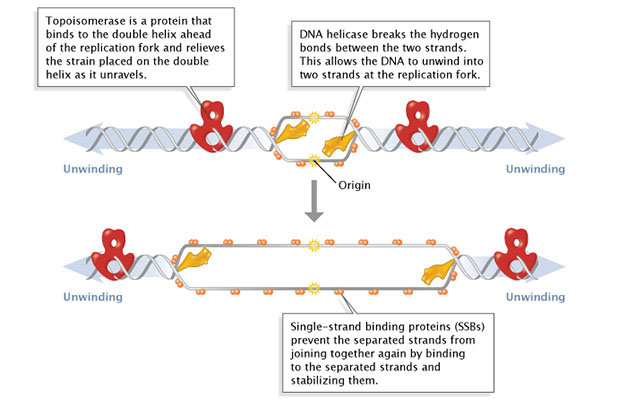
Effigy ane: Facilitation of Deoxyribonucleic acid unwinding.
During DNA replication, several proteins facilitate the unwinding of the Dna double helix into 2 single strands. Topoisomerases (ruby-red) reduce torsional strain caused by the unwinding of the DNA double helix; Deoxyribonucleic acid helicase (yellowish) breaks hydrogen bonds betwixt complementary base of operations-pairs; single-strand binding proteins (SSBs) stabilize the separated strands and prevent them from rejoining.
© 2014 Nature Pedagogy Adapted from Pierce, Benjamin. Genetics: A Conceptual Arroyo, 2nd ed. All rights reserved. ![]()
Equally previously mentioned, the location at which a DNA strand begins to unwind into ii dissever single strands is known as the origin of replication. Equally shown in Figure one, when the double helix unwinds, replication proceeds forth the two single strands at the aforementioned time but in opposite directions (i.east., left to right on one strand, and correct to left on the other). This forms two replication forks that move forth the DNA, replicating as they go.
Primer Synthesis
Primer synthesis marks the beginning of the bodily synthesis of the new Dna molecule. Primers are short stretches of nucleotides (virtually x to 12 bases in length) synthesized by an RNA polymerase enzyme called primase. Primers are required because Deoxyribonucleic acid polymerases, the enzymes responsible for the actual addition of nucleotides to the new DNA strand, can only add deoxyribonucleotides to the 3'-OH group of an existing chain and cannot begin synthesis de novo. Primase, on the other hand, can add together ribonucleotides de novo. Later on, afterwards elongation is complete, the primer is removed and replaced with Dna nucleotides.
Elongation
Finally, elongation--the improver of nucleotides to the new Dna strand--begins after the primer has been added. Synthesis of the growing strand involves adding nucleotides, one by one, in the exact society specified by the original (template) strand. Think that one of the key features of the Watson-Crick Dna model is that adenine is ever paired with thymine and cytosine is always paired with guanine. So, for example, if the original strand reads A-G-C-T, the new strand will read T-C-G-A.
Deoxyribonucleic acid is always synthesized in the v'-to-iii' direction, meaning that nucleotides are added only to the iii' end of the growing strand. As shown in Figure 2, the v'-phosphate group of the new nucleotide binds to the 3'-OH group of the last nucleotide of the growing strand. Scientists have all the same to identify a polymerase that can add together bases to the 5' ends of Deoxyribonucleic acid strands.
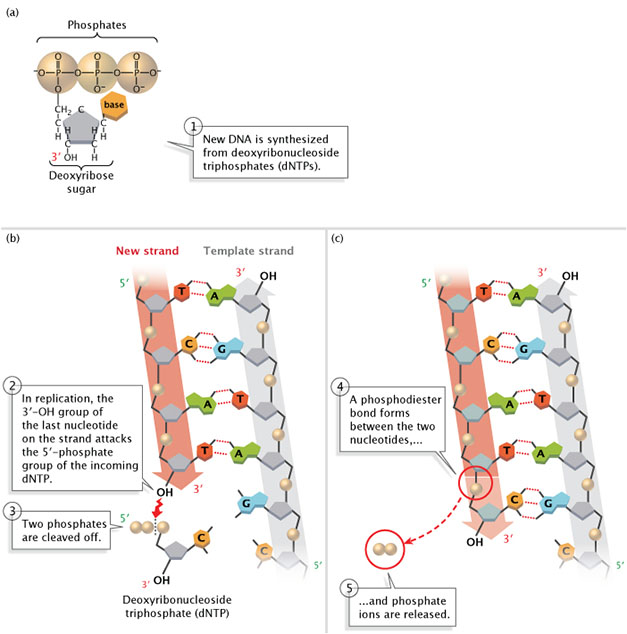
Figure 2: New DNA is synthesized from deoxyribonucleoside triphosphates (dNTPs).
(A) A deoxyribonucleoside triphosphate (dNTP). (B) During Deoxyribonucleic acid replication, the 3'-OH group of the terminal nucleotide on the new strand attacks the 5'-phosphate group of the incoming dNTP. Two phosphates are cleaved off. (C) A phosphodiester bail forms between the ii nucleotides, and phosphate ions are released.
© 2014 Nature Education Adapted from Pierce, Benjamin. Genetics: A Conceptual Approach, 2nd ed. All rights reserved. ![]()
The Discovery of Deoxyribonucleic acid Polymerase
While studying Due east. coli bacteria, enzymologist Arthur Kornberg discovered that Dna polymerases catalyze DNA synthesis. Kornberg'south experiment involved mixing all of the basic "ingredients" necessary for Eastward. coli DNA synthesis in a exam tube, including nucleotides, E. coli extract, and ATP, and then purifying and testing the enzymes involved. Using this method, Kornberg not only discovered Deoxyribonucleic acid polymerases, simply he also performed some of the initial piece of work demonstrating how enzymes add new nucleotides to growing Dna chains (Kornberg, 1959).
Scientists have since identified a total of 5 dissimilar Deoxyribonucleic acid polymerases in E. coli, each with a specialized role. For example, DNA polymerase III does nearly of the elongation piece of work, adding nucleotides 1 by one to the 3' terminate of the new and growing single strand. Other enzymes, including DNA polymerase I and RNase H, are responsible for removing the RNA primer after DNA polymerase III has begun its work, replacing it with Deoxyribonucleic acid nucleotides (Ogawa & Okazaki, 1984). When these enzymes finish, they go out a nick between the section of DNA that was formerly the primer and the elongated section of Dna. Another enzyme called DNA ligase then acts to seal the bond between the two side by side nucleotides.
DNA Polymerase Only Moves in One Direction
Afterward a primer is synthesized on a strand of Dna and the DNA strands unwind, synthesis and elongation tin proceed in only one management. As previously mentioned, Dna polymerase can just add to the 3' cease, so the five' terminate of the primer remains unaltered. Consequently, synthesis proceeds immediately only along the and so-chosen leading strand. This immediate replication is known as continuous replication. The other strand (in the v' direction from the primer) is called the lagging strand, and replication along it is called discontinuous replication. The double helix has to unwind a scrap earlier the synthesis of some other primer can be initiated further up on the lagging strand. Synthesis can then occur from the 3' end of that new primer. Next, the double helix unwinds a scrap more than, and some other spurt of replication gain. As a result, replication forth the lagging strand tin only go on in brusque, discontinuous spurts (Figure 3).
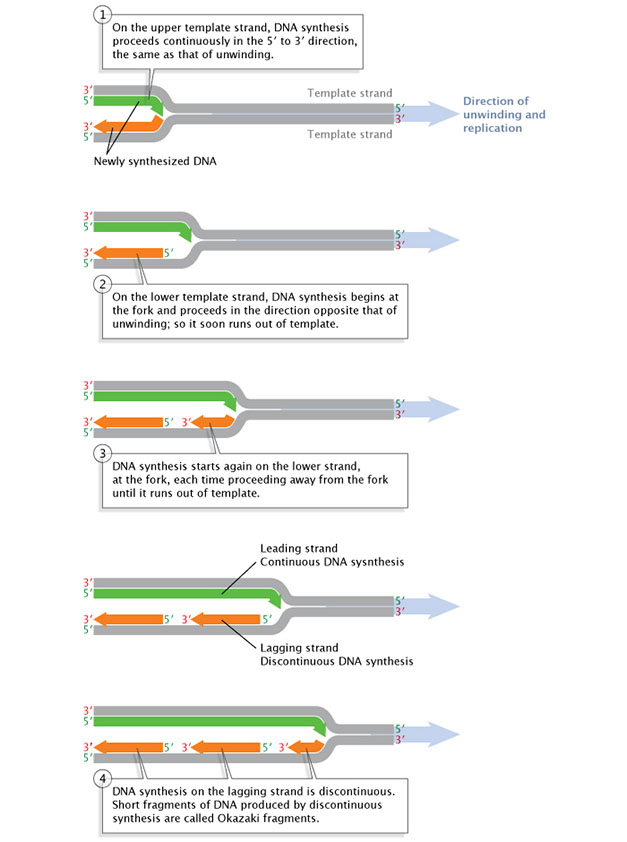
Figure 3: Replication of the leading Dna strand is continuous, while replication along the lagging strand is discontinuous.
Afterward a short length of the DNA has been unwound, synthesis must proceed in the v' to 3' direction; that is, in the direction opposite that of the unwinding.
© 2014 Nature Education Adapted from Pierce, Benjamin. Genetics: A Conceptual Approach, second ed. All rights reserved. ![]()
The fragments of newly synthesized Dna along the lagging strand are called Okazaki fragments, named in honour of their discoverer, Japanese molecular biologist Reiji Okazaki. Okazaki and his colleagues made their discovery by conducting what is known as a pulse-chase experiment, which involved exposing replicating Deoxyribonucleic acid to a short "pulse" of isotope-labeled nucleotides and then varying the length of fourth dimension that the cells would be exposed to nonlabeled nucleotides. This later on period is chosen the "chase" (Okazaki et al., 1968). The labeled nucleotides were incorporated into growing Deoxyribonucleic acid molecules only during the initial few seconds of the pulse; thereafter, just nonlabeled nucleotides were incorporated during the chase. The scientists so centrifuged the newly synthesized DNA and observed that the shorter chases resulted in most of the radioactivity appearing in "slow" DNA. The sedimentation rate was adamant by size: smaller fragments precipitated more slowly than larger fragments because of their lighter weight. As the investigators increased the length of the chases, radioactivity in the "fast" Deoxyribonucleic acid increased with niggling or no increase of radioactivity in the slow DNA. The researchers correctly interpreted these observations to mean that, with short chases, merely very small fragments of Dna were existence synthesized along the lagging strand. Every bit the chases increased in length, giving Deoxyribonucleic acid more time to replicate, the lagging strand fragments started integrating into longer, heavier, more than chop-chop sedimenting Dna strands. Today, scientists know that the Okazaki fragments of bacterial Deoxyribonucleic acid are typically between 1,000 and ii,000 nucleotides long, whereas in eukaryotic cells, they are but about 100 to 200 nucleotides long.
The Challenges of Eukaryotic Replication
Bacterial and eukaryotic cells share many of the same bones features of replication; for instance, initiation requires a primer, elongation is e'er in the v'-to-3' management, and replication is always continuous along the leading strand and discontinuous along the lagging strand. But there are likewise important differences between bacterial and eukaryotic replication, some of which biologists are even so actively researching in an endeavour to better empathize the molecular details. One difference is that eukaryotic replication is characterized by many replication origins (often thousands), not merely one, and the sequences of the replication origins vary widely among species. On the other hand, while the replication origins for bacteria, oriC, vary in length (from about 200 to 1,000 base of operations pairs) and sequence, except among closely related organisms, all bacteria nevertheless have merely a single replication origin (Mackiewicz et al., 2004).
Eukaryotic replication also utilizes a unlike set of Dna polymerase enzymes (e.g., DNA polymerase δ and Deoxyribonucleic acid polymerase ε instead of DNA polymerase Iii). Scientists are still studying the roles of the 13 eukaryotic polymerases discovered to engagement. In addition, in eukaryotes, the Deoxyribonucleic acid template is compacted by the way it winds around proteins called histones. This Deoxyribonucleic acid-histone circuitous, called a nucleosome, poses a unique challenge both for the cell and for scientists investigating the molecular details of eukaryotic replication. What happens to nucleosomes during Deoxyribonucleic acid replication? Scientists know from electron micrograph studies that nucleosome reassembly happens very quickly subsequently replication (the reassembled nucleosomes are visible in the electron micrograph images), but they withal practice not know how this happens (Annunziato, 2005).
As well, whereas bacterial chromosomes are circular, eukaryotic chromosomes are linear. During circular Dna replication, the excised primer is readily replaced past nucleotides, leaving no gap in the newly synthesized DNA. In contrast, in linear DNA replication, there is always a small gap left at the very end of the chromosome because of the lack of a 3'-OH group for replacement nucleotides to bind. (As mentioned, Dna synthesis can go on only in the five'-to-3' direction.) If there were no fashion to make full this gap, the Dna molecule would get shorter and shorter with every generation. Nonetheless, the ends of linear chromosomes—the telomeres—have several properties that preclude this.
DNA replication occurs during the South phase of cell division. In E. coli, this means that the entire genome is replicated in merely twoscore minutes, at a stride of approximately 1,000 nucleotides per second. In eukaryotes, the pace is much slower: about 40 nucleotides per second. The coordination of the poly peptide complexes required for the steps of replication and the speed at which replication must occur in order for cells to divide are impressive, especially considering that enzymes are besides proofreading, which leaves very few errors behind.
Summary
The study of DNA replication started almost as soon as the structure of Dna was elucidated, and it continues to this day. Currently, the stages of initiation, unwinding, primer synthesis, and elongation are understood in the nearly basic sense, but many questions remain unanswered, particularly when it comes to replication of the eukaryotic genome. Scientists take devoted decades to the study of replication, and researchers such as Kornberg and Okazaki accept made a number of of import breakthroughs. Nonetheless, much remains to be learned about replication, including how errors in this process contribute to man affliction.
References and Recommended Reading
Annunziato, A. T. Split decision: What happens to nucleosomes during Deoxyribonucleic acid replication? Journal of Biological Chemistry 280, 12065–12068 (2005)
Bessman, M. J., et al. Enzymatic synthesis of deoxyribonucleic acid. II. Full general properties of the reaction. Journal of Biological Chemistry 233, 171–177 (1958)
Kornberg, A. The biological synthesis of deoxyribonucleic acid. Nobel Lecture, December 11, 1959. (link to transcript)
———. Biological synthesis of deoxyribonucleic acid. Scientific discipline 131, 1503–1508 (1960)
Lehman, I. R., et al. Enzymatic synthesis of deoxyribonucleic acrid. I. Grooming of substrates and fractional purification of an enzyme from Escherichia coli. Journal of Biological Chemistry 233, 163–170 (1958)
Losick, R., & Shapiro, L. Deoxyribonucleic acid replication: Bringing the mount to Mohammed. Science 282, 1430–1431 (1998)
Mackiewicz, P., et al. Where does bacterial replication start? Rules for predicting the oriC region. Nucleic Acids Research 32, 3781–3791 (2004)
Ogawa, T., & Okazaki, T. Role of RNase H in Dna replication revealed past RNase H defective mutants of Escherichia coli. Molecular and General Genetics 193, 231–237 (1984)
Okazaki, R., et al. Mechanism of DNA chain growth. I. Possible discontinuity and unusual secondary structure of newly synthesized chains. Proceedings of the National Academy of Sciences 59, 598–605 (1968)
atkinscallynnusers.blogspot.com
Source: http://www.nature.com/scitable/topicpage/major-molecular-events-of-dna-replication-413#:~:text=DNA%20is%20always%20synthesized%20in,nucleotide%20of%20the%20growing%20strand.
Post a Comment for "Is Dna Read From 5 to 3"